
In 2016, Basketball Reference listed James Harden as a shooting guard. Next year, they switched his position to point guard. It went back to shooting guard in 2018, back to point guard in 2019, and now it’s once again shooting guard in 2020.
The modern NBA is currently in a state of “positional hybridism.” The versatility among players is at an all-time high and it’s a pointless exercise to try to lump them into one of five rigid categories that can’t really describe a player’s skill set. Just look at LeBron James: his Basketball Reference page lists his position as “Power Forward and Point Guard and Small Forward and Shooting Guard.” That’s not very useful!
My goal is to create a new classification of the league’s players through the analysis of data from this season that tells us about a player’s actions on the court. This idea has been done before, usually with somewhere between 20-50 data features. My idea is to take things a bit further. This analysis will include 189 different features — basically every metric that you can think of to describe any player’s playstyle.
I scraped and merged 22 data frames to achieve a data frame with all 189 features used in this project. Including all of that code here would be a little excessive, but you can find all of it in this GitHub repository in the data_collection.py and data_preparation.py files.
After all of that, we can check the shape of df to make sure everything is working as intended:
df.shape
(514, 191)
514 different people have played in an NBA game this season and we have 189 features for our project, which does not include player name and the string indicator for position, so everything’s looking good so far!
Those 514 players include a lot of players who have barely played, though. They don’t offer us enough data to actually conduct any analysis, so let’s set up some minimum requirements to cut down on the observations here. I picked requirements that ensure that our number of observations is still greater than the number of features. I’ll also go ahead and just import everything I need for this analysis in advance.
from sklearn.preprocessing import StandardScaler from sklearn.decomposition import PCA import scipy.cluster.hierarchy as shc from sklearn.cluster import KMeans import matplotlib.pyplot as plt import pandas as pd import numpy as np testdf = df[(df.MPG > 23) & (df.GP > 15)].reset_index(drop=True)
195 players have played at least 15 games while also playing at least 23 minutes per game. Now, let’s get the list of all our features (which is, again, all of our columns minus the player name and the string indicator for position) and get the values of x, the feature values, and y, the player name values.
features = [x for x in df.columns if (x != 'PLAYER_NAME') & (x != 'POSITION')] x = testdf.loc[:, features].values y = testdf.loc[:,['PLAYER_NAME']].values
Many of our features have wildly different ranges and distributions, so let’s be sure to standardize these values before doing anything else.
x = StandardScaler().fit_transform(x)
Now, we can conduct principal component analysis (PCA) to condense these 189 features into just two dimensions. In essence, PCA will create two new features that are made up of the other 189 features in a way that best illustrates the differences between our 195 observations. You can delve far deeper into the linear algebra behind it, but I’ll leave it at that for the purposes of this article. Let’s use PCA and create a new data frame with just three columns: player name along with the two new features.
pca = PCA(n_components=2) principalComponents = pca.fit_transform(x) principalDf = pd.DataFrame(data = principalComponents, columns = ['pc1', 'pc2']) final = pd.concat([principalDf, testdf[['PLAYER_NAME','POSITION','VORP']]], axis=1) final = final[['PLAYER_NAME','POSITION','VORP','pc1','pc2']] final.columns = ['player','pos','vorp','pc1','pc2']
Let’s plot the data:
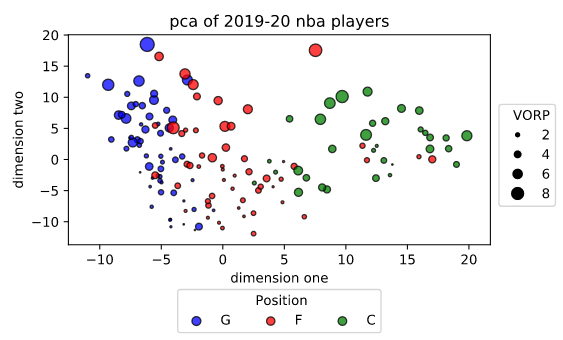
The dimensions are mostly meaningless to us at the moment because we have no idea exactly what they represent. All we know is that they’re made up of our initial 189 features. Looking at this graph, though, we can probably throw out some guesses on what they might represent. The value in dimension one seems to increase as size (assuming centers are bigger than forwards who are bigger than guards) increases, so it’s probably related to rebounding or defense (or both).
The second dimension is probably related to offense because it’s dominated by a group of guards and forwards with high VORP. I’m assuming that includes guys like James Harden, Giannis Antetokounmpo, LeBron James, etc.
We don’t have to guess, though. The following code creates a data frame with the importance of each feature in each dimension:
fimp1 = pd.DataFrame(data={'pc':features,'imp':pca.components_[0]}) # feature importance for first dimension
fimp2 = pd.DataFrame(data={'pc':features,'imp':pca.components_[1]}) # feature importance for second dimensionThis information allows us to plot the features with the biggest positive and negative influence on each dimension. Here’s the plot for the first dimension:
The features with the greatest positive impact on the first dimension relate to setting useful screens, being an efficient pick-and-roll roll man, boxing out, getting rebounds, and scoring off of touches in the paint. These are all skills most important for big men, as expected.
The features with a large negative influence relate to long rebounds, being the pick-and-roll ball handler, dribbling a lot with the ball, shooting threes, driving to the basket, and taking pull-up jumpers. Makes sense — none of these are skills you’d expect from a big man. In a previous article, I specifically found that average rebound distance is negatively correlated with player height. Again, nothing surprising.
Now, let’s take a look at the second dimension.
It seems that the second dimension is largely related to ball-dominance / shot creation.
The extremely negative features include things you’d expect from role players who play off-the-ball: shooting corner threes, a lot of their field goals are assisted, most of their points are from threes, a large percentage of their time is spent without the ball in their hands, they’re often in spot-up situations. Who comes to mind? Probably players like JJ Redick and Duncan Robinson, right?
Meanwhile, the most positive features aren’t even all … well … good things. They are, however, characteristic of a certain type of player. The percent of a team’s turnovers committed by a single player while that player is on the floor and the number of turnovers committed per 36 minutes by that player are both two of the most important features here. Who comes to mind? Ball-dominant players like James Harden and Russell Westbrook, the opposite of players like JJ Redick. Other important features include usage percentage, two-point field goals attempted, and a lot of features related to drawing fouls and getting to the line. I have a feeling James Harden is going to be leading the pack in the first dimension.
Anyway, let’s go back to the scatter plot now that we actually understand our dimensions. It is now possible to create clusters of points based upon their proximity to one another on the plot, which would look something like this:
The issue is that we’re only clustering based upon two dimensions, which combine to explain a limited percentage of the variance in the overall data.
pca.explained_variance_ratio_.cumsum()
array([0.25884462, 0.47164737])
Our transformed two-dimensional data explains just 47.2% of the total variance. That is far from adequate. Instead of using this two-dimensional data for our clustering, we can once again use principal component analysis to find the number of dimensions to reduce our data into that could explain an appropriate percentage of the variance in the data, like 99%. Those dimensions would then be the dimensions we’d use in our cluster analysis in order to create a classification system for NBA players.
Additionally, I think the process of creating a classification system could be improved with a larger number of observations. So, I updated the data collection code from the previous article. The new code does the exact same things as before, but for each of the past seven seasons. Our data now has 1326 observations of 189 features. Much better.
Let’s go ahead and conduct the dimensionality reduction again. Instead of simply reducing our data to two dimensions, we’ll reduce it into the number of dimensions that explain 99% of the variation in the data.
pca = PCA(n_components=0.99) pca.fit(x) principalComponents = pca.transform(x) principalComponents.shape
(1326, 81)
Instead of reducing the 189 initial features into two dimensions, we’ve reduced it to 81 dimensions. We can proceed to validate that these dimensions meet our requirement of explaining 99% of the variance in the data:
pca.explained_variance_ratio_.sum()
0.9900370307594777
Great! We’ve successfully used principal component analysis to reduce the dimensionality of our data and prepare it for a clustering algorithm.
One thing I haven’t really covered, though, is why we’re even bothering going through principal component analysis? Why is dimensionality reduction even important? The answer lies in something called the curse of dimensionality.
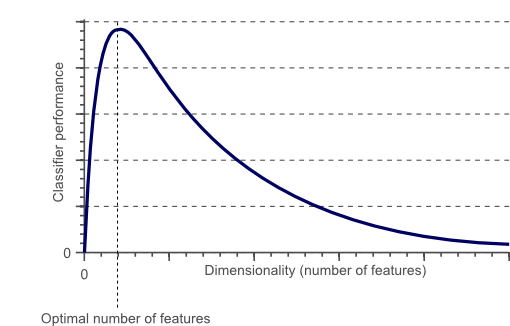
Many clustering algorithms involve a measure of distance between points. At high dimensions, though, these metrics become less meaningful. The average distance between points at high dimensions actually converges, so there is eventually very little difference in distances between different pairs of points. That is obviously not ideal.
In my data collection, I collected basically any feature that I could think of. Therefore, it makes sense to utilize dimensionality reduction to cut down on the ones that aren’t actually descriptive so that the problems that arise with high dimensionality can be avoided. Also, this project does not take much computing power so I went ahead and tested clustering without initially reducing dimensionality and the results were nonsensical.1
I’ll finish the rest of this project in Part II, where we’ll find the optimal number of clusters to split the full data into along with finding an appropriate clustering algorithm for our task.
UPDATE: Part II of this series has been posted at this link.
- Kyrie Irving and Zion Williamson in one cluster, Khris Middleton and Jarrett Allen in another, Isaiah Thomas and Marc Gasol in another, along with Tomas Satoransky and Giannis Antetokounmpo in another. I don’t think any of these pairs of players should belong in the same cluster as one another and the fact that they all do draws some serious questions about the methodology’s reliability.
